|
First, you synthesize the complement, as shown in this image, where dashed blue indicates new synthesis (using regular nucleotides by the way, and DNA polymerase, of course). Now the goal is to make more (a cluster). When you flow the single-stranded library across the flow cell your fragments will base pair with the grafts as shown in the figure below, and you’ll now have your library fragments attached to the flow cell. That’s because those areas are complementary to the P7 and P5 grafts. Note that in the figure above, there are ends that are labelled as P5/P7 graft binding sites. I’m going to call them grafts, and they are the P7 and the P5 grafts. The flow cell (also called chip) has pre-attached short oligos (short sequences of DNA) poking up. More on how many we use per lane above and in another post as as this affects coverage. Each lane can hold (in our work) up to as many as 96 samples = 96 bacterial genomes. They have 8 channels in them, which are called lanes. Flow cells in Illumina look like glass microscope slides. Next, you need to denature the library fragments (break this double-stranded fragment into single strands), and attach the fragments to the flow cell. I’ve drawn a picture below (it’s crude, but hey, it’s mine!) of what those sequences are:Įssentially a library is a sample that contains all of your DNA of interest (in multiple copies), with these adapters (all the coloured sequence) on the ends. Adapters are sequences that belong on the ends. Then those fragments are run through a series of PCR reactions that serve to both amplify the DNA (make more), and place adapaters on them. 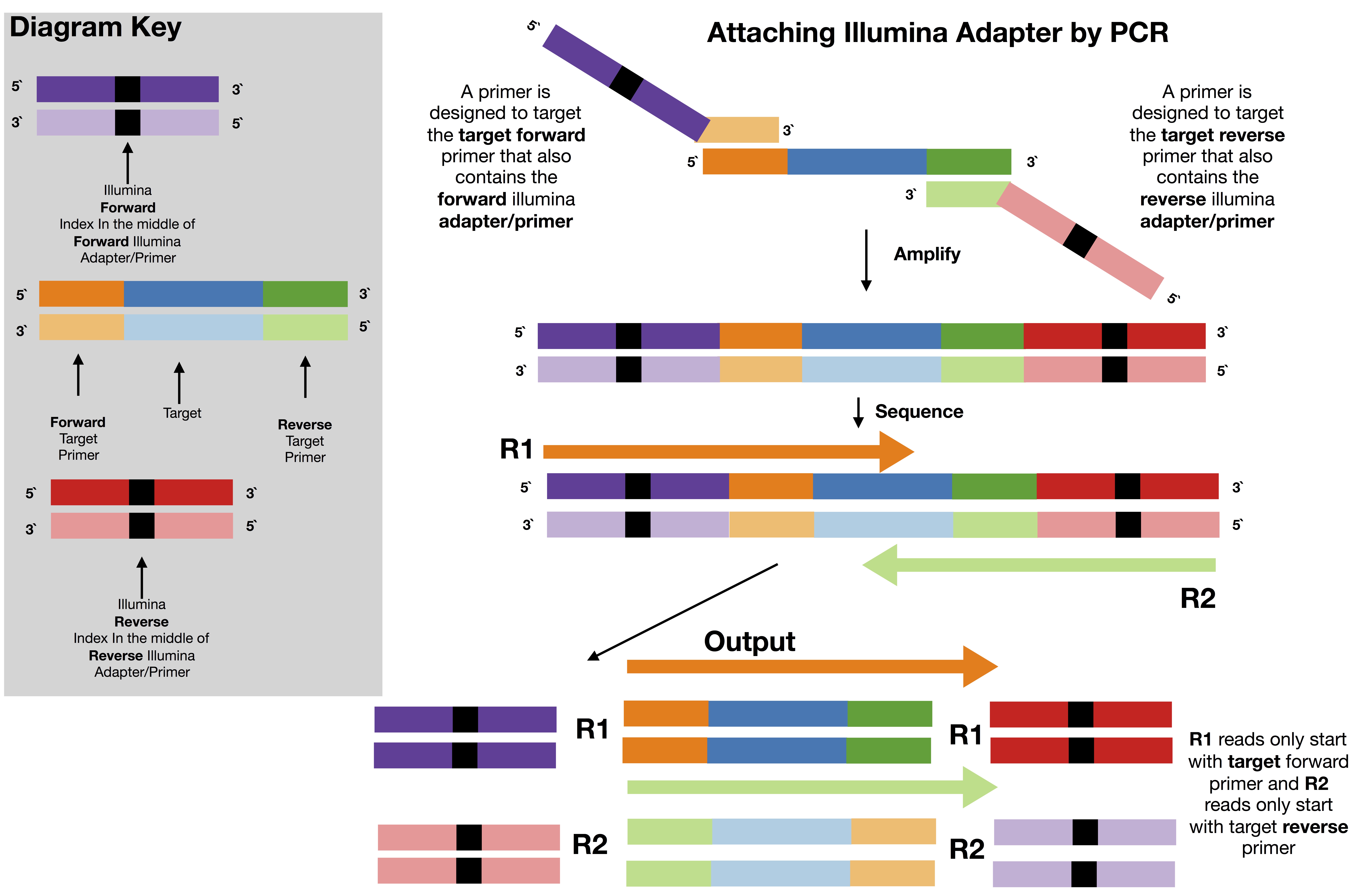
The DNA is fragmented into 350 – 550 bp fragments (I am not sure if this is by sonication, or another method, that’s a question I’m going to ask Steve Simpson). The first thing that happens is library preparation. So what happens once UNH receives our high quality DNA (for us that means PCR amplifiable, looks good on a gel, lots of DNA, verified (we think) to be pure)? More on that and how it affects coverage in another post. We have about 60 samples in a batch and usually have these sequenced on one full lane. 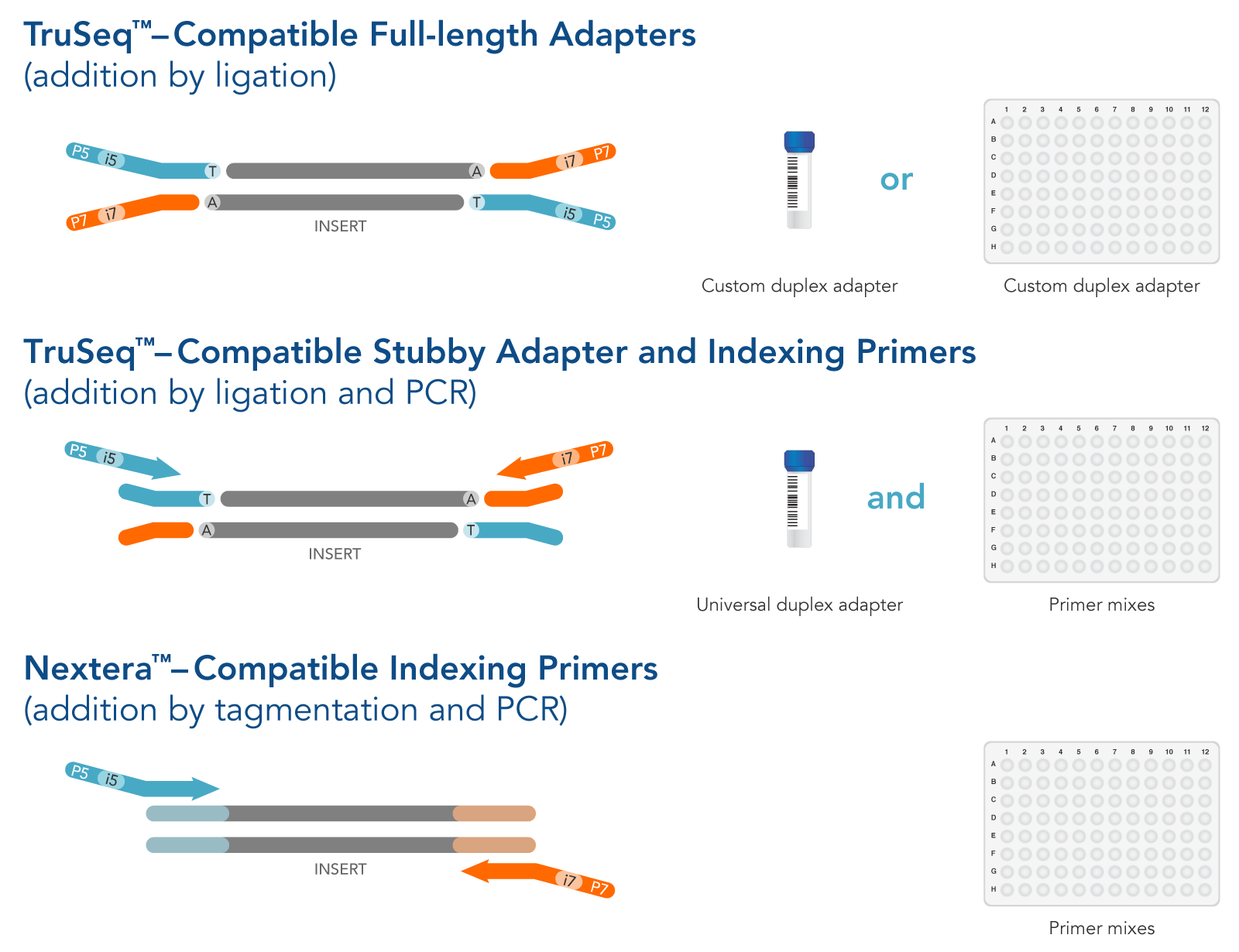
We have 250 bp Paired End (PE) reads done, and our libraries are made by size selecting for fragments not smaller than 350 bp, and not larger than 550 bp. I think I just may finally be cracking this one conceptually…(and I made a set of video lectures about this too, audio isn’t great so consider them beta but “up”).įirst, our samples are sequenced (at the Hubbard Genome Center of UNH) on an Illumina HiSeq 2500 machine. I have struggled with the variety of sources out there that describe Illumina sequencing, none of which seem to be exactly how the samples in my lab are sequenced. I needed a simple, clear explanation of the “for Dummies” variety (I love those books!). For the past year (or so), I have been really struggling to understand the rudiments of how Illumina sequencing works, especially with the concept of “paired ends”.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
